Agena bioscience5/31/2023 
There was no recurrence in any patient with negative ctDNA post-surgery. Post-operative ctDNA samples were available for 13 patients and residual mutations were detected in 10 patients (77%), 3 of whom (30%) had recurrence at median follow up of 20.1 months. 
NRAS and PIK3CA mutations were identified in the ctDNA of 2/3 (67%) and 1/2 (50%) patients. Mutations in NRAS and PIK3CA were identified in 3/27 (11%) and 2/27 (7%) of patients, respectively. 10 patients had a KRAS mutation in their baseline ctDNA that was not detected in their biopsy, with a median MAF of 0.3% (range 0.1-1.3%). This significantly decreased over treatment (0.15% week 3 RT, 0.35% final week RT, 0.1% prior to surgery) (p < 0.05). The median KRAS mutant allele frequency (MAF) in baseline ctDNA samples was 0.9% (range 0.1-2%). Identical mutations were found in baseline ctDNA of 11/15 patients (73%). ResultsĪt least one mutation was identified in 67% (18/27) of baseline biopsy samples. The UltraSEEK™ Colon Panel (Agena Bioscience) was used to track 107 hotspot mutations in serial ctDNA samples. 
DNA from baseline biopsy samples was genotyped for 86 hotspot mutations in BRAF, EGFR, KRAS, NRAS and PIK3CA using the iPLEX™ HS Colon Panel on the MassARRAY® System (Agena Bioscience). MethodsĢ7 LARC patients who were enrolled on the CTRIAL-IE (ICORG) 12-38 TRI-LARC clinical trial (NCT02151019) and received NACRT prior to surgery, had samples for ctDNA analysis taken at baseline pre-NACRT, during week 3 of radiotherapy (RT), during the final week of RT, prior to surgery, and 3-12 months post-surgery. We assessed whether detection and analysis of plasma ctDNA could help monitor tumor evolution during neoadjuvant chemoradiotherapy (NACRT) treatment for LARC. There is limited information on the feasibility and clinical potential of ctDNA analysis in non-metastatic rectal cancer.
0 Comments
Postgres app permission setup5/31/2023  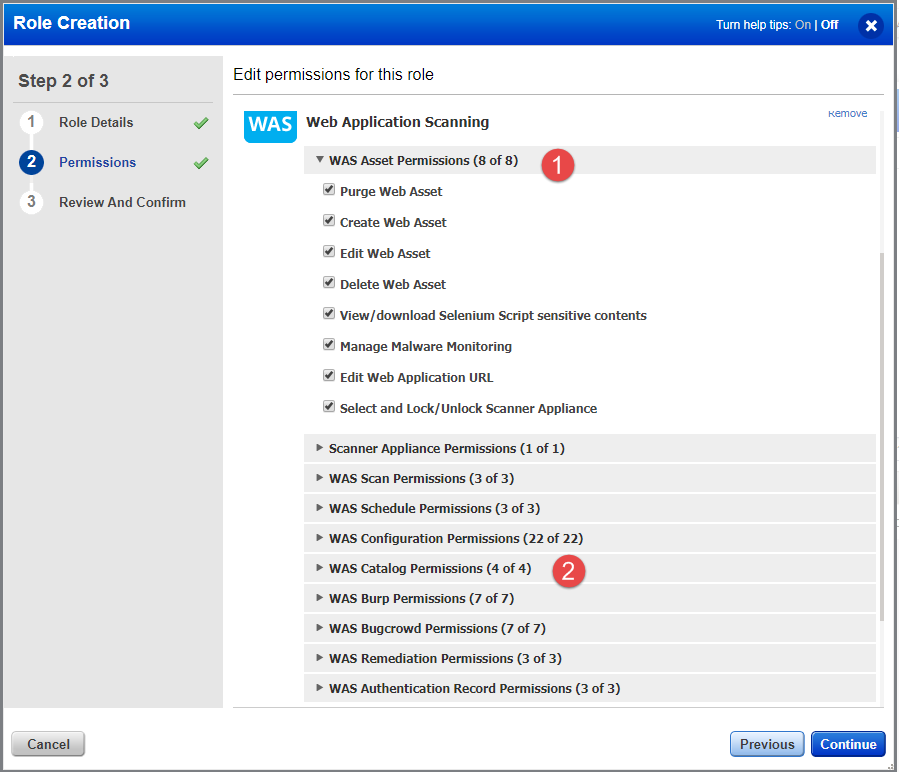
In the Software without restriction, including without limitation the rights Of this software and associated documentation files (the "Software"), to deal The causes and solutions to common errors can be found among the Frequently Asked Questions (FAQ) LicenseĬopyright (c) 2010-2020 Brian Carlson ( is hereby granted, free of charge, to any person obtaining a copy If your change involves breaking backwards compatibility please please point that out in the pull request & we can discuss & plan when and how to release it and what type of documentation or communicate it will require. I will happily accept your pull request if it: If you or your company are benefiting from node-postgres and would like to help keep the project financially sustainable please consider supporting its development. Node-postgres's continued development has been made possible in part by generous finanical support from the community. I try to always announce noteworthy changes & developments with node-postgres on Twitter. You can also follow me if that's your thing. smallest possible snippet of code to reproduce the problem.If you have questions unanswered by the documentation please open an issue pointing out how the documentation was unclear & I will do my best to make it better! If you encounter a bug with the library please open an issue on the GitHub repo. 
The entire list can be found on our wiki. These are some handy modules we've been using over the years to complete the picture. Node-postgres is by design pretty light on abstractions. Bulk import & export with COPY TO/COPY FROM.Named statements with query plan caching.Extensible JS ↔ PostgreSQL data-type coercion.Pure JavaScript client and native libpq bindings share the same API. |
 RSS Feed
RSS Feed
